DC (Direct Current) Resistivity
Model
We assume the following subsurface resistivity distribution with 4 layers:
| Layer # | thickness (m) | |
|---|---|---|
| 1 | 100 | 500 |
| 2 | 200 | 1000 |
| 3 | 20 | 10 |
| 4 | 100 |
The DCModel is defined as :
ρ = log10.([500.0, 1000.0, 10.0, 100.0])
h = [100.0, 200.0, 20.0]
m = DCModel(ρ, h)1D DC Resistivity Model
┌───────┬───────┬────────┬────────┐
│ Layer │ log-ρ │ h │ z │
│ │ [Ωm] │ [m] │ [m] │
├───────┼───────┼────────┼────────┤
│ 1 │ 2.70 │ 100.00 │ 100.00 │
│ 2 │ 3.00 │ 200.00 │ 300.00 │
│ 3 │ 1.00 │ 20.00 │ 320.00 │
│ 4 │ 2.00 │ ∞ │ ∞ │
└───────┴───────┴────────┴────────┘Code for this figure
fig = Figure()
ax = Axis(fig[1, 1]; xlabel="log ρ (log Ωm)", ylabel="depth (m)", backgroundcolor=(
:magenta, 0.05))
plot_model!(ax, m)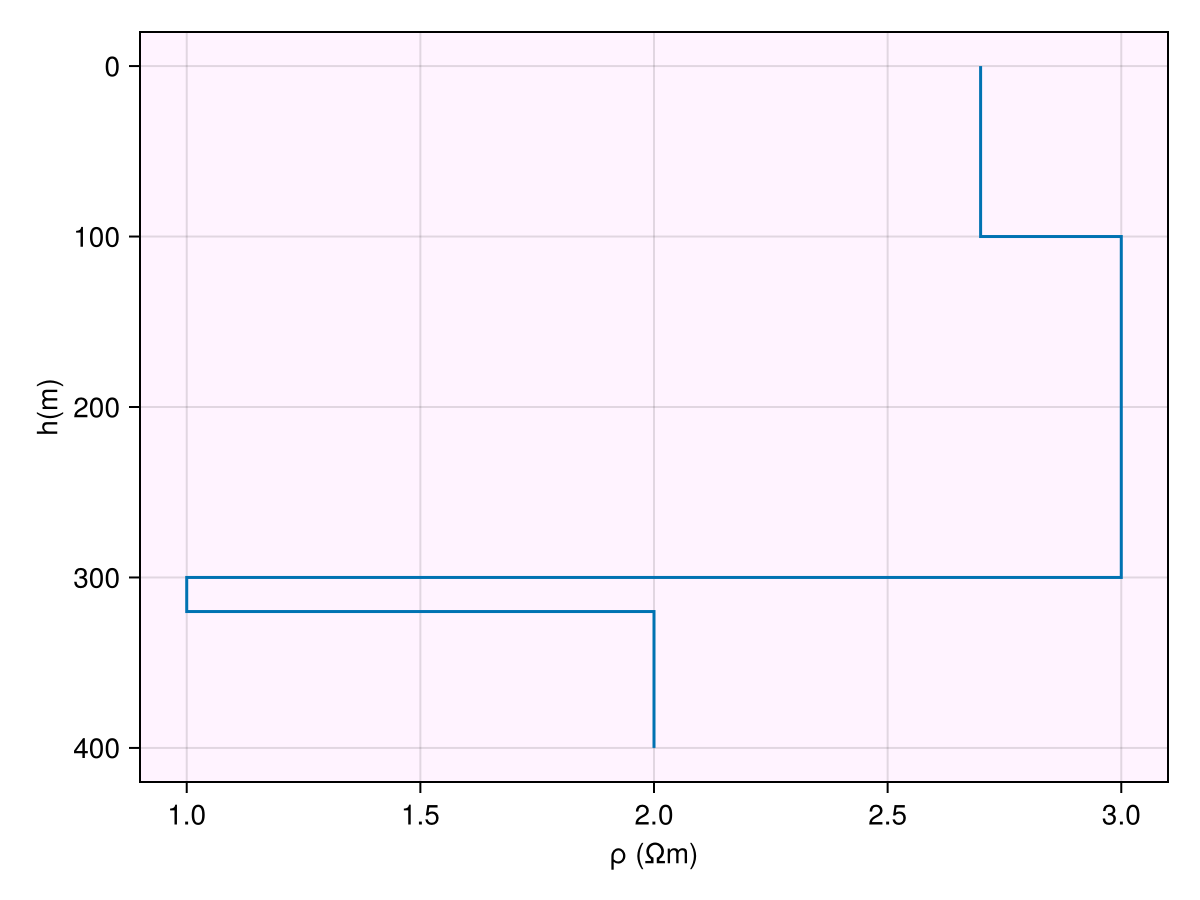
Warn
Always use Float64 or Float32 types while defining the vectors for resistivity and thickness. Using Int will throw an InexactError, e.g. : InexactError: Int64(4193.453970907305)
Response
To obtain the responses, that is apparent resistivity ρₐ, we first need to define the current source and electrodes (receivers) locations. We provide two most common source receiver configurations
PrISM.get_schlumberger_array Function
get_schlumberger_array(a, n)returns electrode spacings for Schlumberger array :
M ←na→ A ←a→ B ←na→ N
Arguments
a: electrode spacing ABn: vector of numbers corresponding to spacing
Usage : TODO
sourcePrISM.get_wenner_array Function
get_wenner_array(a)returns electrode spacings for Wenner array :
M ←a→ A ←a→ B ←a→ N
Arguments
a: vector of electrode spacing AB
Usage
TODO
sourceHere, we demonstrate using the Wenner array for a vertical sounding survey:
locs = get_wenner_array(100:50:2000)(recs = [-50.0 50.0; -75.0 75.0; … ; -975.0 975.0; -1000.0 1000.0], srcs = Float32[-150.0 150.0; -225.0 225.0; … ; -2925.0 2925.0; -3000.0 3000.0])and then call the forward dispatch as usual to get DCResponse:
resp = forward(m, locs)DCResponse{Vector{Float64}}([554.0979075768546, 596.0697919837345, 615.6503568706282, 610.9917723112081, 587.7732852578923, 552.6733686004528, 511.28659870238715, 467.72190083775365, 424.7498699564668, 384.09534170725516 … 114.01883696503232, 112.33447601007096, 110.89607729747706, 109.66998284796625, 108.62494609466799, 107.7312362641314, 106.96272141067718, 106.29731596803161, 105.71409354993084, 105.19324807792805])Code for this figure
fig = Figure() #size = (800, 600))
ax1 = Axis(fig[1, 1]; backgroundcolor=(:magenta, 0.05), aspect=AxisAspect(1))
ab_2 = abs.(locs.srcs[:, 2] .- locs.srcs[:, 1]) ./ 2
plot_response!([ax1], ab_2, resp; plt_type=:scatter)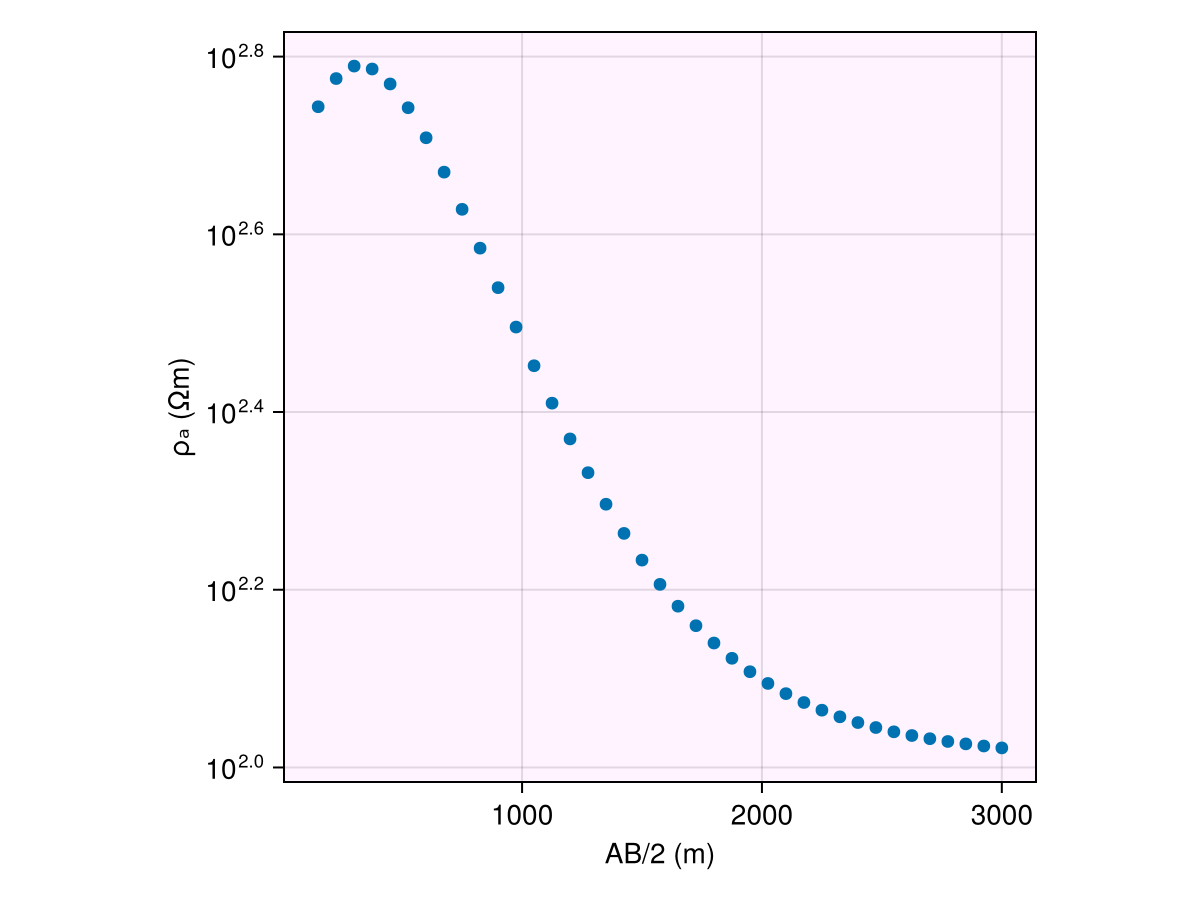
In-place operations
Mutating forward calls are also supported. This leads to no extra allocations while calculating the forward responses. This also speeds up the performance, though the calculations in the well-optimized forward calls are way more expensive than allocations to get a significant boost here. We now call the in-place variant forward! and provide a DCResponse variable to be overwritten :
forward!(resp, m, locs)We can compare the allocations made in the two calls:
@allocated forward!(resp, m, locs)20928@allocated forward(m, locs)21312Benchmarks
While the exact runtimes will vary across processors, the runtimes increase linearly with number of layers.
Benchmark code
n_layers = Int.(exp2.(2:1:6))
locs = get_wenner_array(100:50:2000)
btime_iip = Float64.(zero(n_layers))
btime_oop = Float64.(zero(n_layers))
for i in eachindex(n_layers)
ρ = 2 .* randn(n_layers[i])
h = 100 .* rand(n_layers[i]-1)
m = DCModel(ρ, h)
resp = forward(m, locs)
time_oop = @timed begin
for i in 1:1000
forward(m, locs)
end
end
btime_oop[i] = time_oop.time/1e3
forward!(resp, m, locs)
time_iip = @timed begin
for i in 1:1000
forward!(resp, m, locs)
end
end
btime_iip[i] = time_iip.time/1e3
end
fig = Figure()
ax = Axis(fig[1, 1]; xscale=log2, backgroundcolor=(:magenta, 0.05),
xlabel="no. of layers", ylabel="time (ms)")
lines!(n_layers, btime_oop .* 1e3; label="out-of place forward calls", color=:tomato)
lines!(n_layers, btime_iip .* 1e3; label="in place forward calls",
linestyle=:dash, color=:steelblue3)
Legend(fig[1, 2], ax)┌ Warning: Assignment to `ρ` in soft scope is ambiguous because a global variable by the same name exists: `ρ` will be treated as a new local. Disambiguate by using `local ρ` to suppress this warning or `global ρ` to assign to the existing global variable.
└ @ dc.md:127
┌ Warning: Assignment to `h` in soft scope is ambiguous because a global variable by the same name exists: `h` will be treated as a new local. Disambiguate by using `local h` to suppress this warning or `global h` to assign to the existing global variable.
└ @ dc.md:128
┌ Warning: Assignment to `m` in soft scope is ambiguous because a global variable by the same name exists: `m` will be treated as a new local. Disambiguate by using `local m` to suppress this warning or `global m` to assign to the existing global variable.
└ @ dc.md:129
┌ Warning: Assignment to `resp` in soft scope is ambiguous because a global variable by the same name exists: `resp` will be treated as a new local. Disambiguate by using `local resp` to suppress this warning or `global resp` to assign to the existing global variable.
└ @ dc.md:130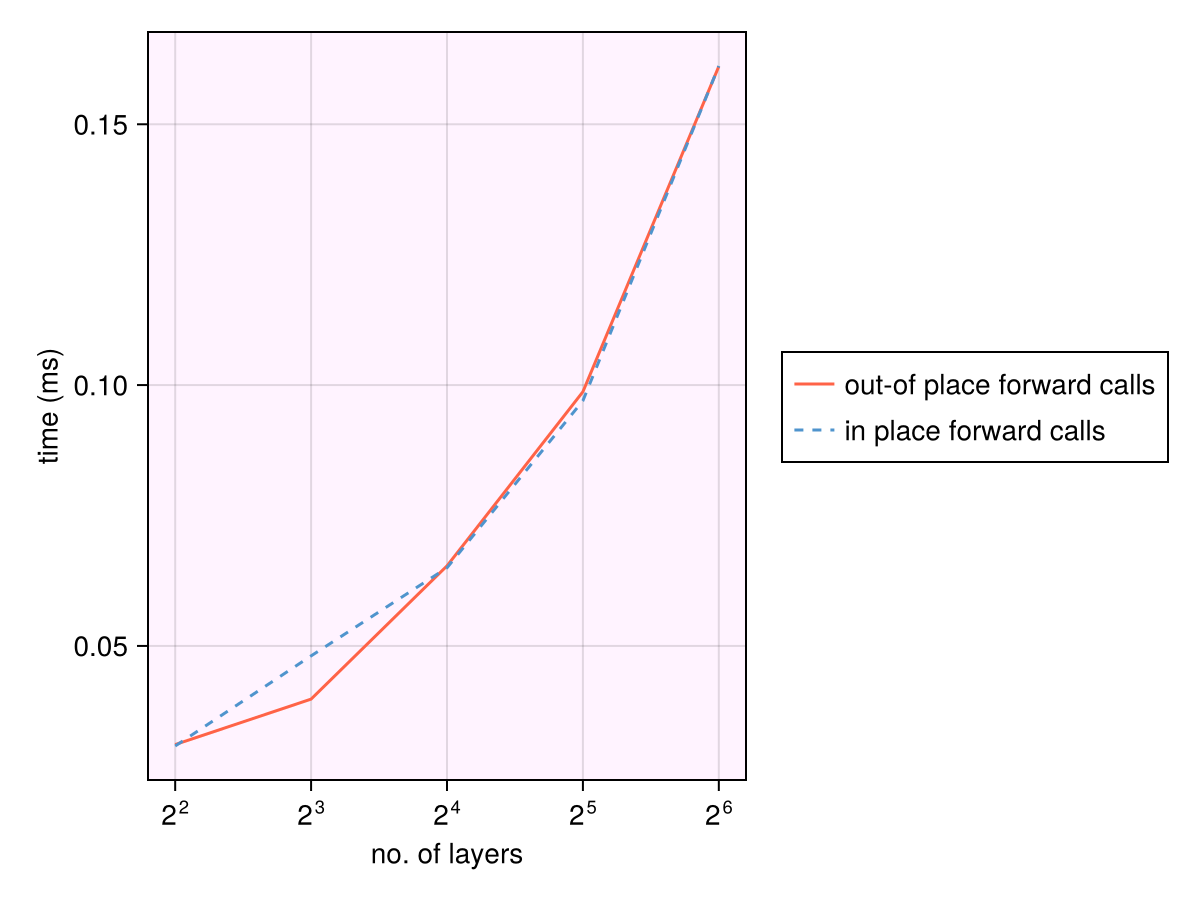
Reproducibility
The above benchmarks were obtained on AMD EPYC 9V74 80-Core Processor