RTO-TKO
RTO-TKO is a stochastic framework introduced in the electromagnetic geophysical community by Blatter et al., 2022 (a and b)
Similar to fixed discretization scheme, the grid sizes do not change. RTO-TKO tries to obtain the samples of the posterior distribution by unrolling the optimization for the misfit function, that is, it perturbs the model space obtained after one iteration before optimizing for the next. It does this iteratively for both the values of the model space and the regularization coefficient in alternative steps.
Therefore, if we have
and
Then for
where
The algorithm was proposed for
Note
Usually, the prior of
is a uniform distribution and we do not have to compute the corresponding log pdf term Implementing the above from scratch might not be trivial because of
being non-invertible, and we do the optimization in the domain defined by . Such a variable will then have a standard normal distribution when
In the following section, we demonstrate RTO-TKO for a 6-layered earth, including the half-space being imaged using the MT method. The prior distribution is composed of 26 layers, with the initial model being a half-space.
Demo
Let's create a synthetic dataset first, with 10% error floors:
m_test = MTModel(log10.([100.0, 10.0, 1000.0]), [1e3, 1e3])
f = 10 .^ range(-4; stop=1, length=25)
ω = vec(2π .* f)
r_obs = forward(m_test, ω)
err_phi = asin(0.02) * 180 / π .* ones(length(ω))
err_appres = 0.1 * r_obs.ρₐ
err_resp = MTResponse(err_appres, err_phi)We don't need to construct the a priori the same way as before. Instead, we need to define the optimiser we'd use. Here, we use Occam, and an initial model composed of half-space of 100
z = collect(0:100:2.5e3)
h = diff(z)
m_rto = MTModel(2 .* ones(length(z)), vec(h))
n_samples = 100
r_cache = rto_cache(
m_rto, [1e-2, 1e4], Occam(), n_samples, n_samples, 1.0, [:ρₐ, :ϕ], false)
mt_chain,
μ_chain = stochastic_inverse(
r_obs, err_resp, ω, r_cache; model_trans_utils=(; m=sigmoid_tf))(MCMC chain (99×26×1 reshape(adjoint(::Matrix{Float64}), 99, 26, 1) with eltype Float64), MCMC chain (99×1×1 Array{Float64, 3}))Note that the chain contains fewer samples than we had asked for. This is because a few unstable samples were filtered out. The obtained mt_chain contains the a posteriori distributions that can be saved using JLD2.jl.
using JLD2
JLD2.@save "file_path.jld2" mt_chainSince RTO-TKO is closely related to Occam, we also include occam results in our figures below. Also, note that we did not require an a priori distribution to obtain samples. However, we would need to create the same to plot our posterior samples. This can just be something that encapsulates the whole prior space (or an envelope of the same), e.g., in the case of magnetotelluric imaging, we know that the resistivity values will always be in [-2, 5] on the log-scale.
modelD = MTModelDistribution(Product([Uniform(-1.0, 5.0) for i in eachindex(z)]), vec(h))Code for this figure
fig = Figure()
ax = Axis(fig[1, 1])
hm = get_kde_image!(ax, mt_chain, modelD; kde_transformation_fn=log10, colormap=:binary,
colorrange=(-3.0, 0.0), trans_utils=(m=no_tf, h=no_tf))
Colorbar(fig[1, 2], hm; label="log pdf")
mean_kws = (; color=:seagreen3, linewidth=2)
std_kws = (; color=:red, linewidth=1.5)
get_mean_std_image!(
ax, mt_chain, modelD; confidence_interval=0.99, trans_utils=(m=no_tf, h=no_tf),
mean_kwargs=mean_kws, std_plus_kwargs=std_kws, std_minus_kwargs=std_kws)
ylims!(ax, [2500, 0])
plot_model!(ax, m_test; color=:black, linestyle=:dash, label="true", linewidth=2)
Legend(fig[2, :], ax; orientation=:horizontal)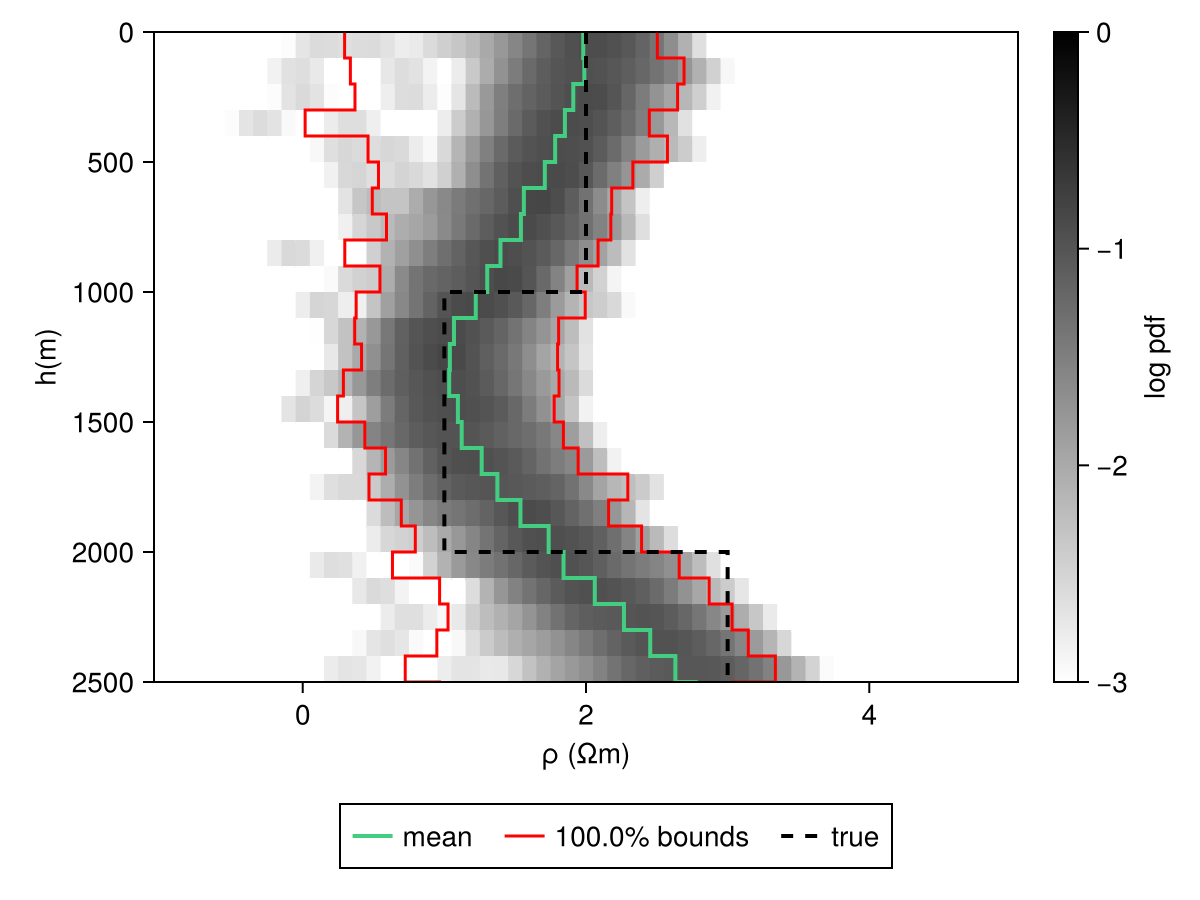
The list of models can then be obtained from chains using
model_list = get_model_list(mt_chain, modelD)We can then easily check the fit of the response curves
fig = Figure()
ax1 = Axis(fig[1, 1])
ax2 = Axis(fig[1, 2])
resp_post = forward(model_list[1], ω);
for i in 1:(length(model_list) > 100 ? 100 : length(model_list))
forward!(resp_post, model_list[i], ω)
plot_response!([ax1, ax2], ω, resp_post; alpha=0.4, color=:gray)
end
plot_response!([ax1, ax2], ω, r_obs; errs=err_resp, plt_type=:errors, whiskerwidth=10)
plot_response!([ax1, ax2], ω, r_obs; plt_type=:scatter, label="true")
ylims!(ax1, exp10.([1.2, 3.1]))
ylims!(ax2, [12, 70])
fig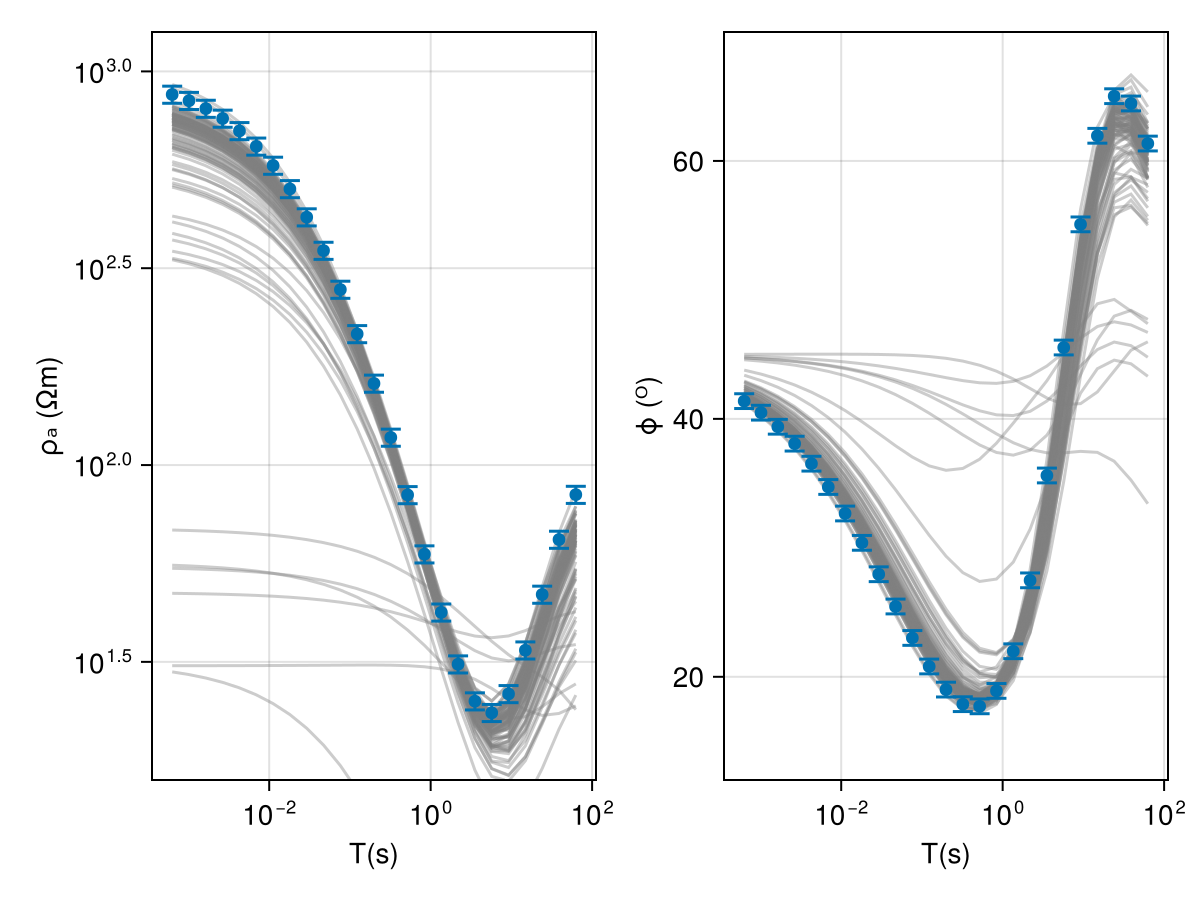
Warning
It is recommended that the samples from RTO-TKO can be filtered out to reject the samples that have a poor fit on the data.