Fixed discretization
Geophysical models generally have fixed discretization. This is mostly because the geophysical inverse problem is already very non-unique, and varying the discretization points together with the model values (e.g. resistivity and velocity) can make the posterior space even wider for model values. While Finite Element schemes have recently stepped in solving the corresponding PDEs, geophysical simulations have traditionally been done using finite differences which do not do well with really non-uniform discretization. Moreover, sensitivity kernels can change drastically if the cell sizes (thickness of the layers) are allowed to vary. We provide the capability to do MCMC inference on such fixed grids.
Let's denote the model parameters, eg., conductivity, by m, and the layer thickness by h. Therefore, in a N-layer case, we will have
such that
and
In the following example, we demonstrate MCMC inversion for a 3-layered earth, including the half-space being imaged using DC resistivity method. The prior distribution assumes all layers have uncorrelated resistivities bounded between
!!! tip Important
The most important thing to be noted here is the specification of the prior distribution, done via:
```julia
modelD = DCModelDistribution(
Product(
[Uniform(-1.0, 5.0) for i in eachindex(z)]
),
vec(h)
);
```
in the example.Demo
Let's create a synthetic dataset first, with 10% error floors:
m_test = DCModel(log10.([100.0, 10.0, 1000.0]), [1e3, 1e3])
locs = get_wenner_array(200:200:5000)
r_obs = forward(m_test, locs)
err_appres = 0.1 * r_obs.ρₐ
err_resp = DCResponse(err_appres)Now, let's define the a priori with fixed grid points
z = collect(0:500:2.5e3)
h = diff(z)
modelD = DCModelDistribution(Product([Uniform(-1.0, 5.0) for i in eachindex(z)]), vec(h))then the likelihood
respD = DCResponseDistribution(normal_dist)Put everything together for MCMC
n_samples = 1000
mcache = mcmc_cache(modelD, respD)
dc_chain = stochastic_inverse(
r_obs, err_resp, locs, mcache, NUTS(), n_samples; progress=false)Chains MCMC chain (1000×20×1 Array{Float64, 3}):
Iterations = 501:1:1500
Number of chains = 1
Samples per chain = 1000
Wall duration = 14.97 seconds
Compute duration = 14.97 seconds
parameters = m[:m][1], m[:m][2], m[:m][3], m[:m][4], m[:m][5], m[:m][6]
internals = n_steps, is_accept, acceptance_rate, log_density, hamiltonian_energy, hamiltonian_energy_error, max_hamiltonian_energy_error, tree_depth, numerical_error, step_size, nom_step_size, logprior, loglikelihood, logjoint
Use `describe(chains)` for summary statistics and quantiles.The obtained dc_chain contains the a posteriori distributions that can be saved using JLD2.jl.
using JLD2
JLD2.@save "file_path.jld2" dc_chainCode for this figure
fig = Figure()
ax = Axis(fig[1, 1])
hm = get_kde_image!(ax, dc_chain, modelD; kde_transformation_fn=log10, colormap=:binary,
colorrange=(-3.0, 0.0), trans_utils=(m=no_tf, h=no_tf))
Colorbar(fig[1, 2], hm; label="log pdf")
mean_kws = (; color=:seagreen3, linewidth=2)
std_kws = (; color=:red, linewidth=1.5)
get_mean_std_image!(
ax, dc_chain, modelD; confidence_interval=0.99, trans_utils=(m=no_tf, h=no_tf),
mean_kwargs=mean_kws, std_plus_kwargs=std_kws, std_minus_kwargs=std_kws)
ylims!(ax, [2500, 0])
plot_model!(ax, m_test; color=:black, linestyle=:dash, label="true", linewidth=2)
Legend(fig[2, :], ax; orientation=:horizontal)Makie.Legend()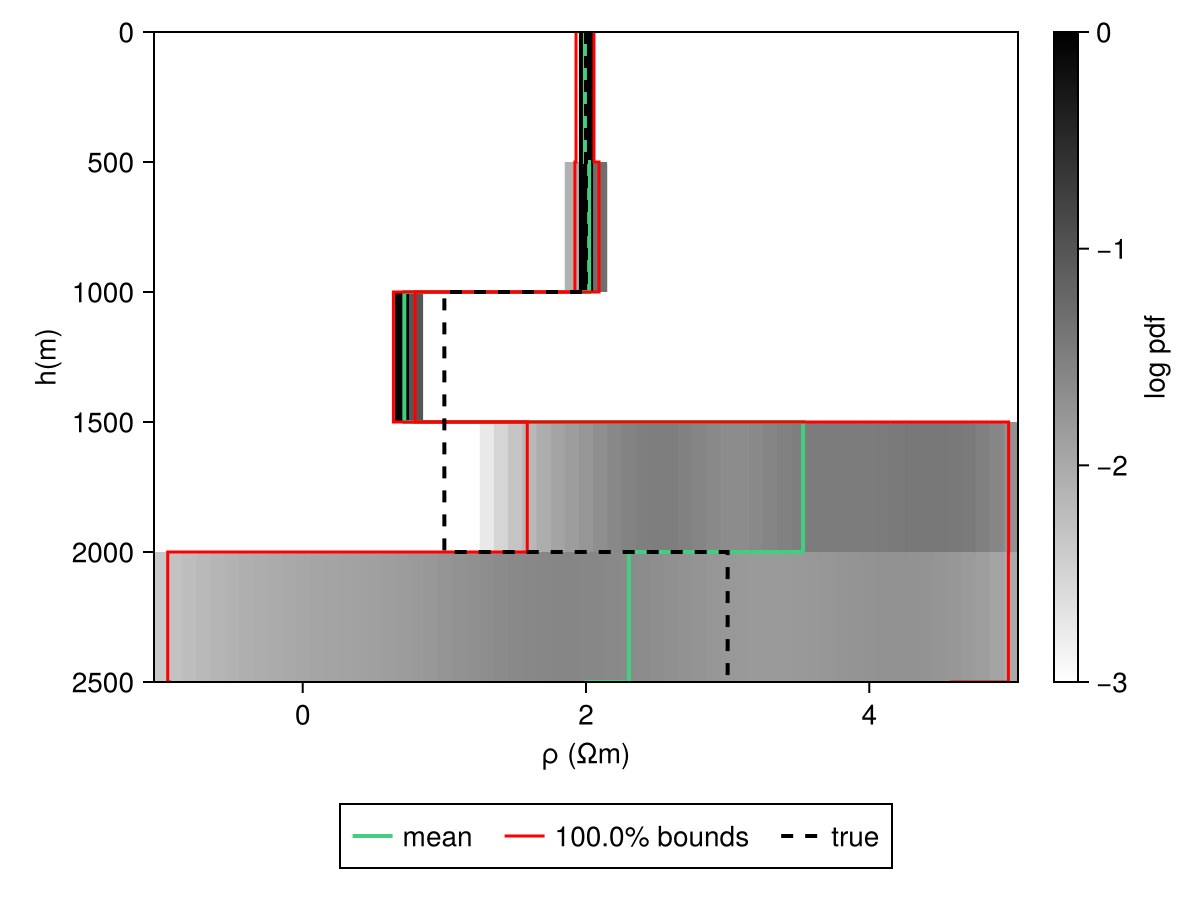
The list of models can then be obtained from chains using, which can then be used to check the fits and perform other diagnostics:
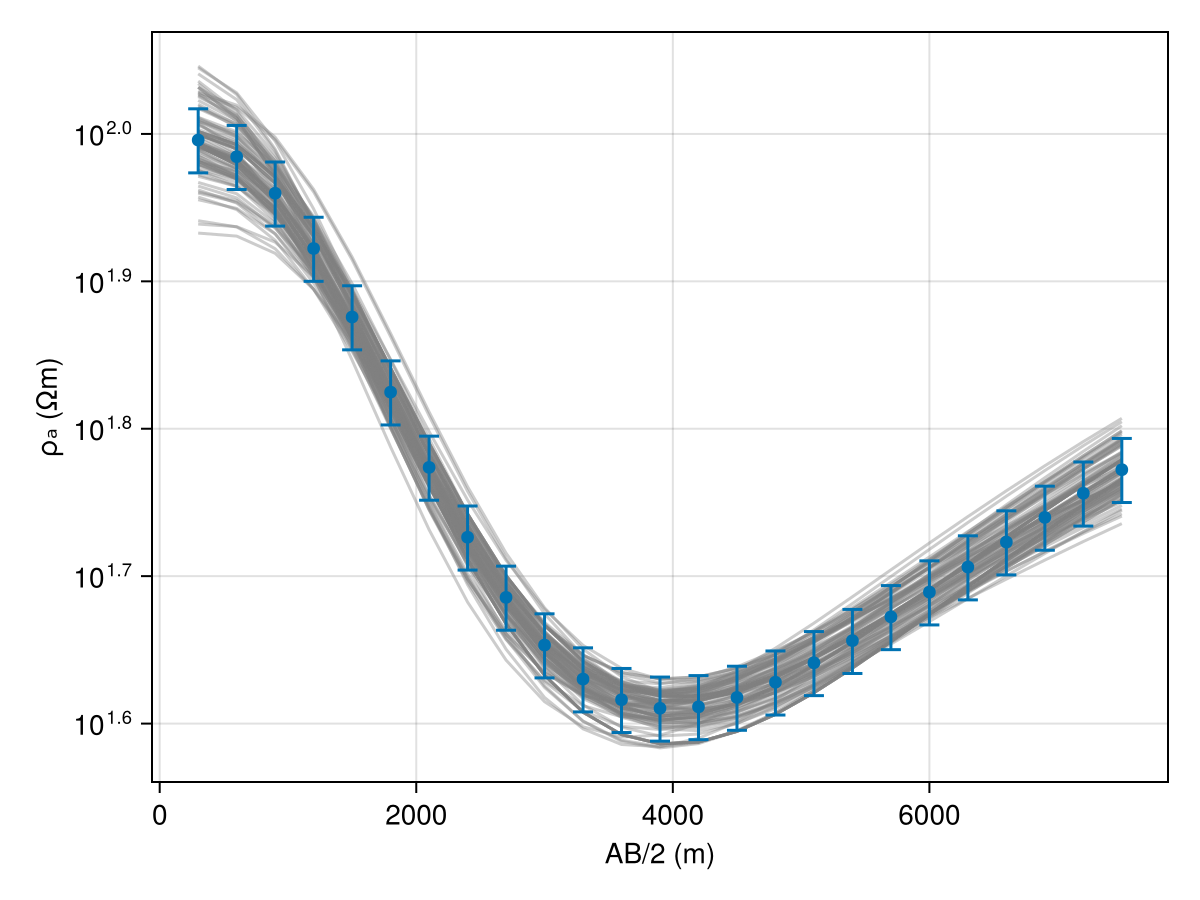
model_list = get_model_list(dc_chain, modelD)Code for this figure
fig = Figure()
ax1 = Axis(fig[1, 1])
ab_2 = abs.(locs.srcs[:, 2] .- locs.srcs[:, 1]) ./ 2
resp_post = forward(model_list[1], locs);
for i in 1:(length(model_list) > 100 ? 100 : length(model_list))
forward!(resp_post, model_list[i], locs)
plot_response!([ax1], ab_2, resp_post; alpha=0.4, color=:gray)
end
plot_response!([ax1], ab_2, r_obs; errs=err_resp, plt_type=:errors, whiskerwidth=10)
plot_response!([ax1], ab_2, r_obs; plt_type=:scatter, label="true")